brief introduction
Using Pandas's pivot method, DF can be rotated and transformed. This paper will explain the secret of pivot in detail.
Using Pivot
pivot is used to reorganize the DF, and use the specified index, columns and values to reconstruct the existing DF.
Take a Pivot example:

Through pivot change, the new DF uses the value in foo as index, the value of bar as columns and zoo as the corresponding value.
Let's take another example of time change:
In [1]: df
Out[1]:
date variable value
0 2000-01-03 A 0.469112
1 2000-01-04 A -0.282863
2 2000-01-05 A -1.509059
3 2000-01-03 B -1.135632
4 2000-01-04 B 1.212112
5 2000-01-05 B -0.173215
6 2000-01-03 C 0.119209
7 2000-01-04 C -1.044236
8 2000-01-05 C -0.861849
9 2000-01-03 D -2.104569
10 2000-01-04 D -0.494929
11 2000-01-05 D 1.071804
In [3]: df.pivot(index='date', columns='variable', values='value') Out[3]: variable A B C D date 2000-01-03 0.469112 -1.135632 0.119209 -2.104569 2000-01-04 -0.282863 1.212112 -1.044236 -0.494929 2000-01-05 -1.509059 -0.173215 -0.861849 1.071804
If the remaining value is more than one column, each column will have a corresponding column value:
In [4]: df['value2'] = df['value'] * 2
In [5]: pivoted = df.pivot(index='date', columns='variable')
In [6]: pivoted
Out[6]:
value value2
variable A B C D A B C D
date
2000-01-03 0.469112 -1.135632 0.119209 -2.104569 0.938225 -2.271265 0.238417 -4.209138
2000-01-04 -0.282863 1.212112 -1.044236 -0.494929 -0.565727 2.424224 -2.088472 -0.989859
2000-01-05 -1.509059 -0.173215 -0.861849 1.071804 -3.018117 -0.346429 -1.723698 2.143608
By selecting value2, you can get the corresponding subset:
In [7]: pivoted['value2'] Out[7]: variable A B C D date 2000-01-03 0.938225 -2.271265 0.238417 -4.209138 2000-01-04 -0.565727 2.424224 -2.088472 -0.989859 2000-01-05 -3.018117 -0.346429 -1.723698 2.143608
Using Stack
Stack is used to convert DF and convert the column into a new internal index.
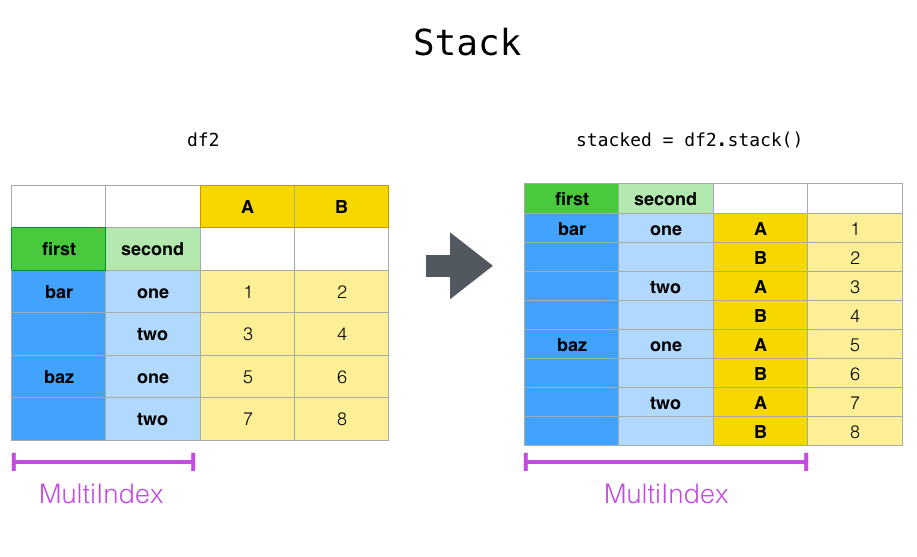
Above, we convert columns A and B to index.
unstack is the reverse operation of stack, which converts the innermost index into the corresponding column.

For example:
In [8]: tuples = list(zip(*[['bar', 'bar', 'baz', 'baz',
...: 'foo', 'foo', 'qux', 'qux'],
...: ['one', 'two', 'one', 'two',
...: 'one', 'two', 'one', 'two']]))
...:
In [9]: index = pd.MultiIndex.from_tuples(tuples, names=['first', 'second'])
In [10]: df = pd.DataFrame(np.random.randn(8, 2), index=index, columns=['A', 'B'])
In [11]: df2 = df[:4]
In [12]: df2
Out[12]:
A B
first second
bar one 0.721555 -0.706771
two -1.039575 0.271860
baz one -0.424972 0.567020
two 0.276232 -1.087401
In [13]: stacked = df2.stack()
In [14]: stacked
Out[14]:
first second
bar one A 0.721555
B -0.706771
two A -1.039575
B 0.271860
baz one A -0.424972
B 0.567020
two A 0.276232
B -1.087401
dtype: float64
By default, unstack is the last index of unstack. We can also specify a specific index value:
In [15]: stacked.unstack()
Out[15]:
A B
first second
bar one 0.721555 -0.706771
two -1.039575 0.271860
baz one -0.424972 0.567020
two 0.276232 -1.087401
In [16]: stacked.unstack(1)
Out[16]:
second one two
first
bar A 0.721555 -1.039575
B -0.706771 0.271860
baz A -0.424972 0.276232
B 0.567020 -1.087401
In [17]: stacked.unstack(0)
Out[17]:
first bar baz
second
one A 0.721555 -0.424972
B -0.706771 0.567020
two A -1.039575 0.276232
B 0.271860 -1.087401

By default, stack will only stack one level, and multiple levels can be passed in:
In [23]: columns = pd.MultiIndex.from_tuples([
....: ('A', 'cat', 'long'), ('B', 'cat', 'long'),
....: ('A', 'dog', 'short'), ('B', 'dog', 'short')],
....: names=['exp', 'animal', 'hair_length']
....: )
....:
In [24]: df = pd.DataFrame(np.random.randn(4, 4), columns=columns)
In [25]: df
Out[25]:
exp A B A B
animal cat cat dog dog
hair_length long long short short
0 1.075770 -0.109050 1.643563 -1.469388
1 0.357021 -0.674600 -1.776904 -0.968914
2 -1.294524 0.413738 0.276662 -0.472035
3 -0.013960 -0.362543 -0.006154 -0.923061
In [26]: df.stack(level=['animal', 'hair_length'])
Out[26]:
exp A B
animal hair_length
0 cat long 1.075770 -0.109050
dog short 1.643563 -1.469388
1 cat long 0.357021 -0.674600
dog short -1.776904 -0.968914
2 cat long -1.294524 0.413738
dog short 0.276662 -0.472035
3 cat long -0.013960 -0.362543
dog short -0.006154 -0.923061
The above is equivalent to:
In [27]: df.stack(level=[1, 2])
Using melt
melt specifies a specific column as a flag variable, and other columns are converted to row data. And placed in two new columns: variable and value.
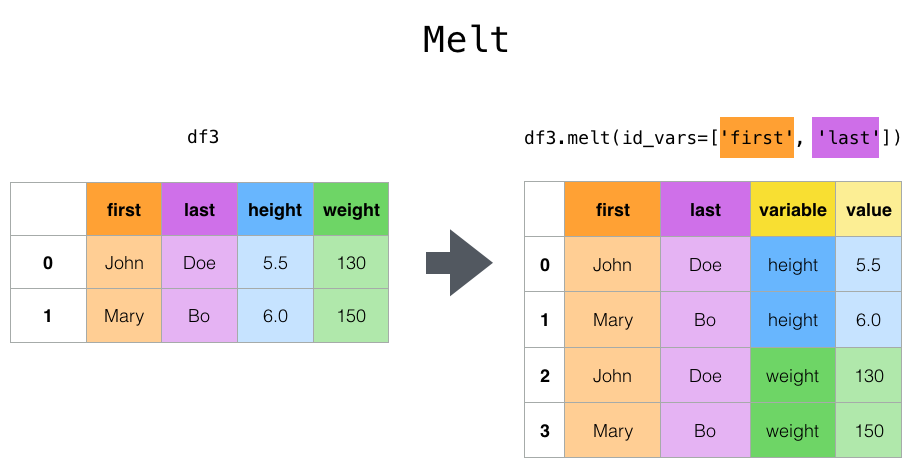
In the above example, we specify two columns, first and last, which are invariant, and height and weight are transformed into row data.
for instance:
In [41]: cheese = pd.DataFrame({'first': ['John', 'Mary'],
....: 'last': ['Doe', 'Bo'],
....: 'height': [5.5, 6.0],
....: 'weight': [130, 150]})
....:
In [42]: cheese
Out[42]:
first last height weight
0 John Doe 5.5 130
1 Mary Bo 6.0 150
In [43]: cheese.melt(id_vars=['first', 'last'])
Out[43]:
first last variable value
0 John Doe height 5.5
1 Mary Bo height 6.0
2 John Doe weight 130.0
3 Mary Bo weight 150.0
In [44]: cheese.melt(id_vars=['first', 'last'], var_name='quantity')
Out[44]:
first last quantity value
0 John Doe height 5.5
1 Mary Bo height 6.0
2 John Doe weight 130.0
3 Mary Bo weight 150.0
Using Pivot tables
Although Pivot can rotate the axis of DF, Pandas also provides pivot_table() can perform numerical statistics while transposing.
pivot_table() receives the following parameters:
data: a df object
values: one or more columns of data to be aggregated.
Index: grouping object of index
Columns: grouping objects for columns
Aggfunc: method of aggregation.
Create a df first:
In [59]: import datetime
In [60]: df = pd.DataFrame({'A': ['one', 'one', 'two', 'three'] * 6,
....: 'B': ['A', 'B', 'C'] * 8,
....: 'C': ['foo', 'foo', 'foo', 'bar', 'bar', 'bar'] * 4,
....: 'D': np.random.randn(24),
....: 'E': np.random.randn(24),
....: 'F': [datetime.datetime(2013, i, 1) for i in range(1, 13)]
....: + [datetime.datetime(2013, i, 15) for i in range(1, 13)]})
....:
In [61]: df
Out[61]:
A B C D E F
0 one A foo 0.341734 -0.317441 2013-01-01
1 one B foo 0.959726 -1.236269 2013-02-01
2 two C foo -1.110336 0.896171 2013-03-01
3 three A bar -0.619976 -0.487602 2013-04-01
4 one B bar 0.149748 -0.082240 2013-05-01
.. ... .. ... ... ... ...
19 three B foo 0.690579 -2.213588 2013-08-15
20 one C foo 0.995761 1.063327 2013-09-15
21 one A bar 2.396780 1.266143 2013-10-15
22 two B bar 0.014871 0.299368 2013-11-15
23 three C bar 3.357427 -0.863838 2013-12-15
[24 rows x 6 columns]
Here are some examples of aggregation:
In [62]: pd.pivot_table(df, values='D', index=['A', 'B'], columns=['C'])
Out[62]:
C bar foo
A B
one A 1.120915 -0.514058
B -0.338421 0.002759
C -0.538846 0.699535
three A -1.181568 NaN
B NaN 0.433512
C 0.588783 NaN
two A NaN 1.000985
B 0.158248 NaN
C NaN 0.176180
In [63]: pd.pivot_table(df, values='D', index=['B'], columns=['A', 'C'], aggfunc=np.sum)
Out[63]:
A one three two
C bar foo bar foo bar foo
B
A 2.241830 -1.028115 -2.363137 NaN NaN 2.001971
B -0.676843 0.005518 NaN 0.867024 0.316495 NaN
C -1.077692 1.399070 1.177566 NaN NaN 0.352360
In [64]: pd.pivot_table(df, values=['D', 'E'], index=['B'], columns=['A', 'C'],
....: aggfunc=np.sum)
....:
Out[64]:
D E
A one three two one three two
C bar foo bar foo bar foo bar foo bar foo bar foo
B
A 2.241830 -1.028115 -2.363137 NaN NaN 2.001971 2.786113 -0.043211 1.922577 NaN NaN 0.128491
B -0.676843 0.005518 NaN 0.867024 0.316495 NaN 1.368280 -1.103384 NaN -2.128743 -0.194294 NaN
C -1.077692 1.399070 1.177566 NaN NaN 0.352360 -1.976883 1.495717 -0.263660 NaN NaN 0.872482
Adding margins=True will add an All column, indicating that All columns are aggregated:
In [69]: df.pivot_table(index=['A', 'B'], columns='C', margins=True, aggfunc=np.std)
Out[69]:
D E
C bar foo All bar foo All
A B
one A 1.804346 1.210272 1.569879 0.179483 0.418374 0.858005
B 0.690376 1.353355 0.898998 1.083825 0.968138 1.101401
C 0.273641 0.418926 0.771139 1.689271 0.446140 1.422136
three A 0.794212 NaN 0.794212 2.049040 NaN 2.049040
B NaN 0.363548 0.363548 NaN 1.625237 1.625237
C 3.915454 NaN 3.915454 1.035215 NaN 1.035215
two A NaN 0.442998 0.442998 NaN 0.447104 0.447104
B 0.202765 NaN 0.202765 0.560757 NaN 0.560757
C NaN 1.819408 1.819408 NaN 0.650439 0.650439
All 1.556686 0.952552 1.246608 1.250924 0.899904 1.059389
Using crosstab
Crosstab is used to count the number of occurrences of elements in a table.
In [70]: foo, bar, dull, shiny, one, two = 'foo', 'bar', 'dull', 'shiny', 'one', 'two' In [71]: a = np.array([foo, foo, bar, bar, foo, foo], dtype=object) In [72]: b = np.array([one, one, two, one, two, one], dtype=object) In [73]: c = np.array([dull, dull, shiny, dull, dull, shiny], dtype=object) In [74]: pd.crosstab(a, [b, c], rownames=['a'], colnames=['b', 'c']) Out[74]: b one two c dull shiny dull shiny a bar 1 0 0 1 foo 2 1 1 0
The crosstab can receive two Series:
In [75]: df = pd.DataFrame({'A': [1, 2, 2, 2, 2], 'B': [3, 3, 4, 4, 4],
....: 'C': [1, 1, np.nan, 1, 1]})
....:
In [76]: df
Out[76]:
A B C
0 1 3 1.0
1 2 3 1.0
2 2 4 NaN
3 2 4 1.0
4 2 4 1.0
In [77]: pd.crosstab(df['A'], df['B'])
Out[77]:
B 3 4
A
1 1 0
2 1 3
You can also use normalize to specify the scale value:
In [82]: pd.crosstab(df['A'], df['B'], normalize=True) Out[82]: B 3 4 A 1 0.2 0.0 2 0.2 0.6
You can also normalize rows or columns:
In [83]: pd.crosstab(df['A'], df['B'], normalize='columns') Out[83]: B 3 4 A 1 0.5 0.0 2 0.5 1.0
You can specify the aggregation method:
In [84]: pd.crosstab(df['A'], df['B'], values=df['C'], aggfunc=np.sum) Out[84]: B 3 4 A 1 1.0 NaN 2 1.0 2.0
get_dummies
get_dummies can convert a column in DF into a combination of 0 and 1 of k columns:
df = pd.DataFrame({'key': list('bbacab'), 'data1': range(6)})
df
Out[9]:
data1 key
0 0 b
1 1 b
2 2 a
3 3 c
4 4 a
5 5 b
pd.get_dummies(df['key'])
Out[10]:
a b c
0 0 1 0
1 0 1 0
2 1 0 0
3 0 0 1
4 1 0 0
5 0 1 0
get_dummies and cut can be combined to count the elements in the range:
In [95]: values = np.random.randn(10)
In [96]: values
Out[96]:
array([ 0.4082, -1.0481, -0.0257, -0.9884, 0.0941, 1.2627, 1.29 ,
0.0824, -0.0558, 0.5366])
In [97]: bins = [0, 0.2, 0.4, 0.6, 0.8, 1]
In [98]: pd.get_dummies(pd.cut(values, bins))
Out[98]:
(0.0, 0.2] (0.2, 0.4] (0.4, 0.6] (0.6, 0.8] (0.8, 1.0]
0 0 0 1 0 0
1 0 0 0 0 0
2 0 0 0 0 0
3 0 0 0 0 0
4 1 0 0 0 0
5 0 0 0 0 0
6 0 0 0 0 0
7 1 0 0 0 0
8 0 0 0 0 0
9 0 0 1 0 0
get_dummies can also accept a DF parameter:
In [99]: df = pd.DataFrame({'A': ['a', 'b', 'a'], 'B': ['c', 'c', 'b'],
....: 'C': [1, 2, 3]})
....:
In [100]: pd.get_dummies(df)
Out[100]:
C A_a A_b B_b B_c
0 1 1 0 0 1
1 2 0 1 0 1
2 3 1 0 1 0
This article has been included in http://www.flydean.com/05-python-pandas-reshaping-pivot/
The most popular interpretation, the most profound dry goods, the most concise tutorial, and many tips you don't know are waiting for you to find!
Welcome to my official account: "those things in procedure", understand technology, know you better!