Parameter setting
There are often problems in XGB classification
There are parameters to adjust the sample imbalance
scale_pos_weight
,
Usually, we enter the ratio of negative sample size to positive sample size in the parameter
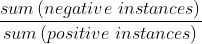
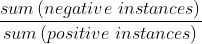

Classification case
Create unbalanced dataset
import numpy as np
import xgboost as xgb
import matplotlib.pyplot as plt
from sklearn.datasets import make_blobs
from sklearn.model_selection import train_test_split as TTS
from sklearn.metrics import accuracy_score as accuracy, recall_score as recall, roc_auc_score as auc
class_1 = 500 # Category 1 has 500 samples
class_2 = 50 # Category 2 has only 50
centers = [[0.0, 0.0], [2.0, 2.0]] # Set the center of two categories
clusters_std = [1.5, 0.5] # Set the variance of the two categories. Generally speaking, the category with large sample size will be more loose
X, y = make_blobs(n_samples=[class_1, class_2],
centers=centers,
cluster_std=clusters_std,
random_state=0, shuffle=False)
Xtrain, Xtest, Ytrain, Ytest = TTS(X,y,test_size=0.3,random_state=420)
(y == 1).sum() / y.shape[0]
XGBoost modeling
dtrain = xgb.DMatrix(Xtrain,Ytrain)
dtest = xgb.DMatrix(Xtest,Ytest)
param= {'silent':True,'objective':'binary:logistic',"eta":0.1,"scale_pos_weight":1}
num_round = 100
bst = xgb.train(param, dtrain, num_round)
preds = bst.predict(dtest)
The above preds returns the classification probability of the sample. The predicted category of the sample can be generated by setting the threshold below.
#Set your own threshold ypred = preds.copy() ypred[preds > 0.5] = 1 ypred[ypred != 1] = 0
Parameter adjustment evaluation
# Parameter setting
scale_pos_weight = [1,5,10]
names = ["negative vs positive: 1",
"negative vs positive: 5",
"negative vs positive: 10"]
for name,i in zip(names,scale_pos_weight):
param={'silent':True,'objective':'binary:logistic',"eta":0.1,"scale_pos_weight":i,'verbosity':0}
clf = xgb.train(param, dtrain, num_round)
preds = clf.predict(dtest)
ypred = preds.copy()
ypred[preds > 0.5] = 1
ypred[ypred != 1] = 0
print(name)
print("\tAccuracy:{}".format(accuracy(Ytest,ypred)))
print("\tRecall:{}".format(recall(Ytest,ypred)))
print("\tAUC:{}".format(auc(Ytest,preds)))
print("\t")
# Different thresholds can also be used
for name,i in zip(names,scale_pos_weight):
for thres in [0.3,0.5,0.7,0.9]:
param= {'silent':True,'objective':'binary:logistic',"eta":0.1,"scale_pos_weight":i,'verbosity':0}
clf = xgb.train(param, dtrain, num_round)
preds = clf.predict(dtest)
ypred = preds.copy()
ypred[preds > thres] = 1
ypred[ypred != 1] = 0
print("{},thresholds:{}".format(name,thres))
print("\tAccuracy:{}".format(accuracy(Ytest,ypred)))
print("\tRecall:{}".format(recall(Ytest,ypred)))
print("\tAUC:{}".format(auc(Ytest,preds)))
print("\t")
Result analysis and suggestions
- The ratio of negative sample size to positive sample size in the original data set is 10:1, and the model parameter scale is compared in turn_ pos_ After the value of weight is 1, 5 and 10, it is found that the accuracy, AUC area and recall rate of the model on the test set are improved, indicating that this parameter does make the XGBoost model have a good processing ability for unbalanced data.
- In essence, scale_ pos_ The weight parameter is adjusted by adjusting the predicted probability value. You can observe how the probability is affected by viewing the results returned by bst.predict(Xtest). Therefore, when we only care about whether the predicted results are accurate and whether the AUC area or recall rate is good enough, we can use scale_pos_weight parameter to help us.
- However, xgboost has many other functions besides classification and regression. In some fields that need to use accurate probability (such as ranking), we hope to maintain the original appearance of probability and improve the effect of the model. At this time, we can't use scale_pos_weight to help us.
- If we only care about the overall performance of the model, we use AUC as the model evaluation index and scale_pos_weight to deal with the sample imbalance problem. If we care about the probability of predicting the correct probability, we can't adjust the scale_pos_weight to reduce the impact of sample imbalance. In this case, we need to consider another parameter: max_delta_step. This parameter is called "the single maximum increment allowed in the weight estimation of the tree", which can be considered as a parameter affecting the estimation. If we are dealing with the problem of sample imbalance and are very concerned about getting the correct prediction probability, we can set max_ delta_ The step parameter is a finite number (such as 1) to help convergence. max_ delta_ The step parameter is usually not used. It is the only purpose of this parameter when the sample imbalance problem under the second classification is.